Ricardo Pereira - Hybridization and Speciation
Hybridization is common throughout the tree of life, and plays profound impacts for patterns of biodiversity, leading to species collapse, rapid adaptation or completion of species formation. I explore the diversity of natural hybridization systems, including invertebrates, amphibians, and mammals, in both field- and lab-based approaches, to provide insights on the process of species formation. In particular my research tackles the following fundamental questions: How do genes interact with each other or with the environment?; How do these interactions result in fitness changes in individuals?; How does fitness affect species splitting or fusion?

Polytypic species as windows into the continuum of species formation
The tempo at which reproductive isolation evolves between diverging lineages and how hybridization may change that dynamics have remained two fundamental question in evolutionary biology. To tackle these questions, I focus on species complexes showing multiple hybrid zones between lineages at continuous stages of divergence. By developing population genetic methods for non-model organisms that often lack reference genomes 1, we have shown that a continuum of genetic interactions can persist over long evolutionary time 2,3 and that hybridization has a heterogeneous effect in different parts of the genome 4,5.
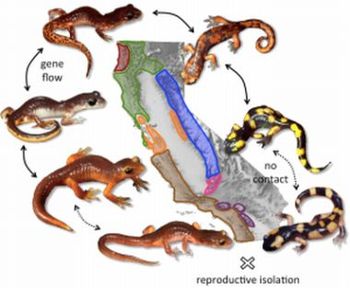
For example, subspecies of the salamander Ensatina eschscholtzii have been diverging around the inhospitable Californian Central Valley for ~8Mya. Using population genetic methods I’ve demonstrated that genetic interactions upon secondary contact reflect a large continuum of possibilities expected for ring species, from unrestricted gene flow, to stable hybrid zones, and finally to overlap with full reproductive isolation 2. Taking advantage of this continuum of divergence, I have shown that the frequency of hybrids is predicted by the level of nuclear genetic divergence, but not by ecological or geographic isolation 3. top
Experimental methods as a link between genotype and organismal fitness
Genomic methods provide valuable lists of candidate genes that potentially underlie species formation. Yet, the link between genes and the phenotypes affecting individual fitness remains obscure, particularly in non-model organisms. In my research, I tackle this question by raising pure populations and their hybrids in common garden 6 and assess fitness in controlled laboratorial conditions 7,8.

Populations of the copepods have been diverging along the coast of North America. Using experimental hybridization we have we found that enzymatic pathways related to the mitochondria are dysfunctional in hybrids 6, suggesting that cyto-nuclear incompatibilities are important at early stages of speciation 9. Using split-broods, we demonstrated that thermal adaptation has a strong genetic basis 8. Using recombinant inbred lines we have shown that a few generations of recombination can purge out incompatibilities while allowing for gene combinations with additive effects on thermal fitness 7, potentially leading to new ecological niches.
Towards an integration between field- and lab-based approaches
Despite the underlying idiosyncrasies of species formation, two patterns are general across species, termed as the two rules of speciation. First, ‘Haldane’s rule’ states that whenever a sex is absent, rare or sterile in hybrids, that sex is usually the heterogametic sex. Second, the ‘large X-effect’ states that the X or Z chromosome has a disproportionately high impact on hybrid fitness compared to autosomes. These patterns have been extensively described on experimental hybrids in model organisms that do not hybridize in nature.

The hybrid zone of the grasshopper Chorthippus parallelus obeys to the two rules of speciation, while interacting in an old natural hybrid zone, providing a remarkable system to test if these rules actually contribute for species formation in nature. Currently, we are using transcriptomics to assess patterns of gene flow across the hybrid zone formed after the last glaciation. Using experimental hybrids, we will be able to test candidate genes functionally, and provide links between phenotypes, genes, and gene flow between emerging species.top
Selected publications:
- da Fonseca, R. R. et al. Next-generation biology: Sequencing and data analysis approaches for non-model organisms. Marine Genomics 1–11 (2016). doi:10.1016/j.margen.2016.04.012
- Pereira, R. J. & Wake, D. B. Genetic leakage after adaptive and nonadaptive divergence in the Ensatina eschscholtzii ring species. Evolution 63, 2288–2301 (2009).
- Pereira, R. J., Monahan, W. B. & Wake, D. B. Predictors for reproductive isolation in a ring species complex following genetic and ecological divergence. BMC Evol Biol 11, 194 (2011).
- Pereira, R. J., Martínez-Solano, I. & Buckley, D. Hybridization during altitudinal range shifts: nuclear introgression leads to extensive cyto-nuclear discordance in the fire salamander. Molecular Ecology 25, 1551–1565 (2016).
- Carneiro, M. et al. The genomic architecture of population divergence between subspecies of the european rabbit. Plos Genetics 10, e1003519 (2014).
- Barreto, F. S., Pereira, R. J. & Burton, R. S. Hybrid Dysfunction and Physiological Compensation in Gene Expression. Molecular Biology and Evolution 32, 613–622 (2015).
- Pereira, R. J., Barreto, F. S. & Burton, R. S. Ecological novelty by hybridization: experimental evidence for increased thermal tolerance by transgressive segregation in Tigriopus californicus. Evolution 68, 204–215 (2014).
- Pereira, R. J., Sasaki, M. C. & Burton, R. S. Adaptation to a latitudinal thermal gradient within a widespread copepod species: the contributions of genetic divergence and phenotypic plasticity. Proceedings of the Royal Society of London B. in press
- Burton, R. S., Pereira, R. J. & Barreto, F. S. Cytonuclear Genomic Interactions and Hybrid Breakdown. Annu Rev Ecol Evol S 44, 281–302 (2013).

